Huilin Li Laboratory
Cryo-EM, DNA Replication and Repair, and Protein Glycosylation and Homeostasis
In recent years, advances in cryo-electron microscopy (cryo-EM) have revolutionized the study of life’s most basic components. Scientists can now image molecules and molecular complexes more precisely than ever before, providing mechanistic insight into the biological building blocks that play key roles in normal health and in disease. The ability to see these tiny molecules gives scientists a powerful tool when designing new therapeutic strategies for some of humanity’s greatest health threats.
The laboratory of Dr. Huilin Li leverages state-of-the-art cryo-EM technology to several ends, including uncovering the molecular machinery behind eukaryotic DNA replication and repair, which can contribute to tumorigenesis when it goes awry; investigating how proteins are modified by various sugars (glycosylation); how chaperones assist protein folding and transport them to their intracellular or extracellular destinations; and how damaged or unwanted proteins are degraded and recycled.
Recent News & Press
Learn More
A sticky situation: How a molecular ‘adhesive’ may be the key to making an antibiotic more effective

Cryo-EM reveals never-before-seen look at the ‘proofreading’ proteins that safeguard DNA
When it comes to DNA replication, humans and baker’s yeast are more alike than different
Our Impact
We're raising thousands to save millions
We’re turning hope into action for the millions of people around the world affected by diseases like cancer and Parkinson’s. Find out how you can help us make a difference.
- 122 peer-reviewed papers published in 2024, 63 of which were in high-impact journals
- 15 VAI-SU2C Epigenetics Dream Team clinical trials launched to date
- 10 clinical trials co-funded by VAI & Cure Parkinson's (out of 41 total International Linked Clinical Trials Program trials)
Huilin Li, Ph.D.
Chair and Professor, Department of Structural Biology; Ralph and Grace Hauenstein Endowed Chair in Structural Biology
Areas of Expertise
Structural biology, cryo-electron microscopy, biophysics, DNA replication, membrane protein biochemistry
Biography
Huilin Li, Ph.D., is an internationally recognized structural biologist with more than 20 years of experience in cryo-electron microscopy (cryo-EM). His current work focuses on the eukaryotic DNA replication, the mycobacterial proteasome system, and the protein folding and protein glycosylation.
Dr. Li earned his Ph.D. in electron crystallography from the University of Science and Technology Beijing, where he trained with the late Prof. K.H. Kuo. He then completed postdoctoral research in the labs of Dr. Bing Jap and Dr. Kenneth Downing at Lawrence-Berkeley National Laboratory, where he studied membrane channels and microtubule structure by cryo-EM. From there, he joined Brookhaven National Laboratory as an associate biophysicist, rising through the ranks to attain a tenured position. In 2010, he joined Stony Brook University as a professor in the Department of Biochemistry and Cell Biology while also maintaining a summer appointment at Brookhaven. He is now a professor and chair of Van Andel Institute’s Department of Structural Biology.
Eukaryotic DNA Replication
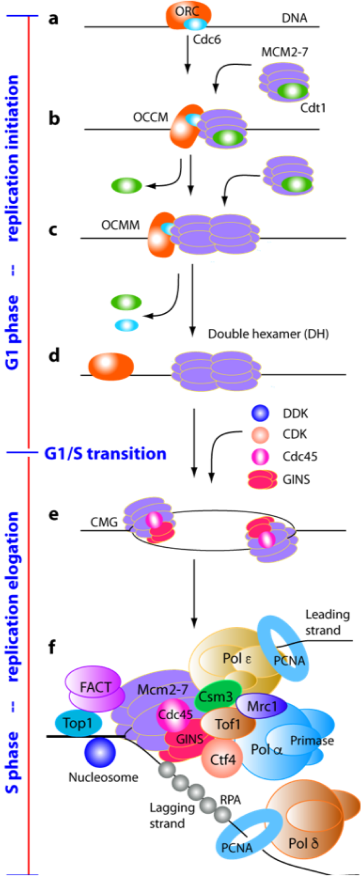
Eukaryotic chromosomal replication initiation is an intricate process that requires the coordinated and tightly regulated action of numerous molecular machines. Failure to ensure once-only replication initiation per cell cycle can result in uncontrolled proliferation and genomic instability, two hallmarks of tumorigenesis. The origin recognition complex (ORC) is a six-protein machine conserved in all eukaryotes. Yeast ORC constitutively binds to and marks the replication origin throughout the cell cycle. Licensing of the DNA replication origin starts when the critical cell division cycle protein Cdc6p binds to ORC in the G1 phase of the cell cycle.
In collaboration with the labs of Dr. Christian Speck and Dr. Bruce Stillman, we have mapped the architecture of ORC (Chen et al. 2008, Proc Natl Acad Sci U S A), captured several key intermediates during origin activation and Mcm2-7 hexamer recruitment on to DNA (Sun et al. 2013, Nat Struct Mol Biol; Sun et al. 2014, Genes Dev), and elucidated how Cdc6 completes the ORC ring and activates it to load the replicative helicase (Yuan et al. 2017, Nat Struct Mol Biol; Feng et al. 2021, Nat Commun). In 2017, we showed how the two Mcm2-7 hexamers come together on the origin DNA to form the pre-replication complex — the Mcm2-7 double-hexamer (Noguchi et al. 2017, Proc Natl Acad Sci U S A).
In the S phase of the cell cycle, the active CMG helicase works with the leading strand polymerase epsilon, the lagging strand polymerase delta, and the primase-polymerase alpha to synthesize new DNA. In collaboration with Dr. Michael O’Donnell’s lab, we have determined the holoenzyme structures of the leading strand DNA polymerase epsilon (Yuan et . 2020, Nat Commun), the lagging strand DNA polymerase delta (Zheng et al. 2020, Proc Natl Acad Sci U S A), and polymerase alpha-primase complex (Yuan et al. 2023, Nat Commun). We found Ctf4 trimer can coordinate two CMG helicases and one Pol alpha-primase into a replication factory core (Yuan et al. 2019, eLife).
Replicative polymerases are tethered to DNA by the PCNA sliding clamp. The PCNA is a topologically closed ring that needs to be cracked open to encircle DNA, and this is accomplished by the clamp loaders. We discovered that the canonical clamp loader RFC has evolved unique structural features to recognize both the 3′- and 5′-DNA ends to load PCNA onto gaps for lagging strand DNA synthesis and for DNA repair (Zheng et al. 2022, eLife). PCNA loading to the leading strand DNA is uniquely challenging because the 3’ DNA junction may be preoccupied by Polε recruited to DNA during origin activation. We have discovered how the yeast Ctf18-RFC binds to PCNA, DNA and Polε, leading to PCNA ring opening and lateral transfer of DNA from Polε to PCNA (Yuan et al. 2024, Science). We have also determined a series of PCNA loading intermediates of the human CTF18-RFC (He et al. 2024, Proc Natl Acad Sci U S A).
The PCNA ring needs to be taken off or unloaded from DNA upon completion of DNA synthesis. The PCNA unloading mechanism is least understood. We have discovered that both yeast Elg1-RFC and human ATAD5-RFC have evolved “locking loops” and a “plug domain” to prevent DNA binding, explaining why the Elg1/ATAD5-RFC are an exclusive PCNA unloader (Wang et al. 2024, Nat Struct Mol Biol).
In addition to PCNA, eukaryotes have evolved another DNA encircling ring called 9-1-1, which is a heterotrimer that functions in DNA damage checkpoint signaling. The 9-1-1 ring is loaded to the 5′ DNA by Rad24-RFC. We have demonstrated how Rad24-RFC loads the 9-1-1 checkpoint clamp ring onto the recessed 5′-end by binding a 5′-DNA at an external surface site and threading the 3′-ssDNA into the chamber and into 9-1-1 (Zheng et al. 2022, Nat Struct Mol Biol), how Rad24-RFC binds to a 10-nt and a 5-nt gap DNA, explaining why Rad24-RFC is unable to melt DNA ends and Rad24-RFC’s preference for a preexisting gap of >5 nt and suggesting a role of the 9-1-1 in gap repair (Zheng et al. 2023, Cell Rep).
Our work has advanced the field of eukaryotic DNA replication (Li et al. 2018, BioEssay; O’Donnell et al. 2018, Nat Struct Mol Biol; Yuan et al. 2020, Biochem J).
Related publications
Chen Z, Speck C, Wendel P, Tang C, Stillman B, Li H. 2008. The architecture of the DNA replication origin recognition complex in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 105(30):10326–10331.
Feng X, Noguchi Y, Barbon M, Stillman B, Speck C, Li H. 2021. The structure of ORC-Cdc6 on an origin DNA reveals the mechanism of ORC activation by the replication initiator Cdc6. Nat Commun 12(1):3883.
Georgescu R, Yuan Z, Bai L, de Luna Almeida Santos R, Sun J, Zhang D, Yurieva O, Li H*, O’Donnell ME*. 2017. Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation. Proc Natl Acad Sci U S A 114(5):E697–E706.
He Q, Wang F, Yao NY, O’Donnell ME*, Li H*. 2024. Structures of the human leading strand Polε–PCNA holoenzyme. Nat Commun 15(7847).
*Co-corresponding authors
He Q, Wang F, O’Donnell ME, Li H. 2024. Cryo-EM reveals a nearly complete PCNA loading process and unique features of the human alternative clamp loader CTF18-RFC. Proc Nat Acad Sci U S A 121(18):e2319727121.
Li H, O’Donnell ME. 2018. The eukaryotic CMG helicase at the replication fork: Emerging architecture reveals an unexpected mechanism. BioEssays 40(3).
Noguchi Y, Yuan Z, Bai L, Schneider S, Zhao G, Stillman B, Speck C, Li H. 2017. Cryo-EM structure of Mcm2-7 double hexamer on DNA suggests a lagging-strand DNA extrusion model. Proc Natl Acad Sci U S A 114(45):E9529–E9538.
O’Donnell ME, Li H. 2018. The ring-shaped hexameric helicases that function at DNA replication forks. Nat Struct Mol Biol 25(2):122–130.
Sun J, Evrin C, Samel S, Fernadez-Cid A, Riera A, Kawakami H, Zech J, Stillman B, Speck C, Li H. 2013. Cryo-EM structure of a helicase loading intermediate containing ORC-Cdc6-Cdt1-MCM2-7 bound to DNA. Nat Struct Mol Biol 20(8):944–951.
Sun J, Fernandez-Cid A, Riera A, Tognetti S, Yuan Z, Stillman B, Speck C, Li H. 2014. Structural and mechanistic insights into licensing of DNA replication. Genes Dev 28:2291–2303.
Sun J, Shi Y, Georgescu RE, Yuan Z, Chait BT, Li H*, O’Donnell ME*. 2015. The architecture of a eukaryotic replisome. Nat Struct Mol Biol 22(12):976–982.
Wang F, He Q, Yao NY, O’Donnell ME, Li H. 2024. The human ATAD5 has evolved unique structural elements to function exclusively as a PCNA unloader. Nat Struct Mol Biol.
Yuan Z, Bai L, Sun J, Georgescu R, O’Donnell ME, Li H. 2016. Structure of the eukaryotic replicative CMG helicase suggests a pumpjack motion for translocation. Nat Struct Mol Biol 23(3):217–224.
Yuan Z, Georgescu R, Yao NY, Yurieva O, O’Donnell ME, Li H. 2024. Mechanism of PCNA loading by Ctf18-RFC for leading-strand DNA synthesis. Science 385(6708).
Yuan Z, Georgescu R, Li H*, O’Donnell ME*. 2023. Molecular choreography of primer synthesis by the eukaryotic Pol α-primase. Nat Commun 14(1):3697.
*Co-corresponding authors
Yuan Z, Georgescu R, Santos RLA, Zhang D, Bai L, Yao NY, Zhao G, O’Donnell ME, Li H. 2019. Ctf4 organizes sister replisomes and Pol α into a replication factory. eLife e47405.
Yuan Z, Georgescu R, Schauer GD, O’Donnell ME, Li H. 2020. Structure of the polymerase ε holoenzyme and atomic model of the leading strand replisome. Nat Commun11(1):3156.
Yuan Z, Li H. 2020. Molecular mechanisms of eukaryotic origin initiation, replication fork progression, and chromatin maintenance. Biochem J 477(18):3499–3525.
Yuan Z, Riera A, Bai L, Sun J, Nandi S, Spanos C, Chen ZA, Barbon M, Rappsilber J, Stillman B, Speck C, Li H. 2017. Structural basis of MCM2-7 replicative helicase loading by ORC-Cdc6 and Cdt1. Nat Struct Mol Biol 24(3):316–324.
Zheng F, Georgescu RE, Li H, O’Donnell ME. 2020. Structure of eukaryotic DNA polymerase δ bound to the PCNA clamp while encircling DNA. Proc Natl Acad Sci U S A117(48):30344–30353.
Zheng F, Georgescu R, Yao NY, Li H, O’Donnell ME. 2022. Cryo-EM structures reveal that RFC recognizes both the 3′- and 5′-DNA ends to load PCNA onto gaps for DNA repair. eLife 11:e77469.
Zheng F, Yao NY, Georgescu RE, Li H*, O’Donnell ME*. 2024. Structure of the PCNA unloader Elg1-RFC. Sci Adv 10(9).
*Co-corresponding authors
Zheng F, Georgescu RE, Yao NY, O’Donnell ME, Li H. 2023. Structures of 9-1-1 DNA checkpoint clamp loading at gaps from start to finish and ramification on biology. Cell Rep 42(7):112694.
Protein Glycosylation and Trafficking
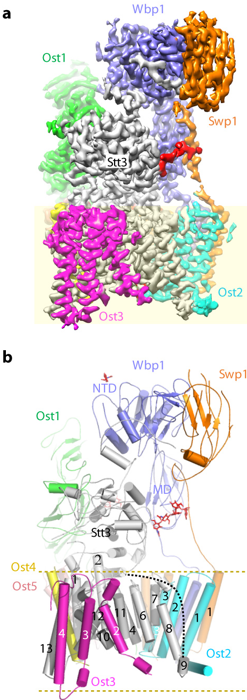
Proteins can be modified with sugars either via an N-linkage on asparagine or via an O-linkage on the OH of a serine or threonine. N-glycosylation in eukaryotes is catalyzed by an ER-embedded eight-protein membrane complex oligosaccharyltransferase (OST). Our earlier work mapped the subunit arrangement of the OST complex (Li et al. 2008, Structure), and showed that OST binds to ribosome at a location near the nascent peptide exit tunnel (Harada et al. 2009, Proc Natl Acad Sci U S A). Recently, we determined the atomic model of the OST, revealing the functions for many of the subunits and, unexpectedly, several phospholipids mediating subunit-subunit interaction in the transmembrane region (Bai et al. 2018, Nature). Protein O-mannosyltransferase forms either a heterodimer or a homodimer, and associates with translocon in the ER membrane to perform co-translational protein O-mannosylation. We found the Pmt1-Pmt2 heterodimer has the GT-C fold and share structural homology with OST, confirming that protein O-mannosylation and protein N-glycosylation are evolutionarily related (Bai et al. 2019, Nat Struct Mol Biol).
O-glucosylation is a novel type of O-glycosylation. We recently studied enzymes that add a glucose-xylose-xylose trisaccharide on Notch receptor. We solved the structure of Rumi that adds the first glucose to Notch and found that Rumi recognizes a six-amino-acid sequence on many of the 36 EGF motifs of Notch and a hydrophobic patch (Yu et al. 2016, Nat Chem Biol). Our study suggested that mutated forms of Rumi could be the cause of cancers because these mutant enzymes are unable to modify Notch. This observation links Rumi defects with cancers. XXYLT1 is a retaining glycosyltransferase that adds the third sugar of the trisaccharide — a xylose — to Notch. The chemical mechanism for retaining enzymes such as XXYLT1 had been controversial in the field for several decades. Our structural studies unambiguously supported the SNi-like retaining mechanism and disproved the double displacement mechanism (Yu et al. 2015, Nat Chem Biol). We further solved the structure of an EGF motif covalently modified by a full O-glucose trisaccharide (Takeuchi et al. 2017, J Biol Chem). The structure reveals that the glycan fills up a surface groove of the EGF with multiple contacts with the protein, providing a chemical basis for the stabilizing effects of the glycans.
The core of heparin sulfate is a copolymer of two alternating sugars GlcA and GlcNAc. This copolymer core is synthesized by exostosin-1 and exostosin-2, which form an obligatory heterodimer (EXT1-2). By solving a series structures of the enzyme complexed with donors and acceptors, we have recently shown that each subunit contains one activity: the EXT1 GT-B domain contains the β1,4GlcA transferase domain and the EXT2 GT-A domain the α1,4GlcNAc transferase domain (Li et al. 2023, Nat Chem Biol). Because the two catalytic sites are far apart (90 Å), we suggested that HS is synthesized by a dissociative process.
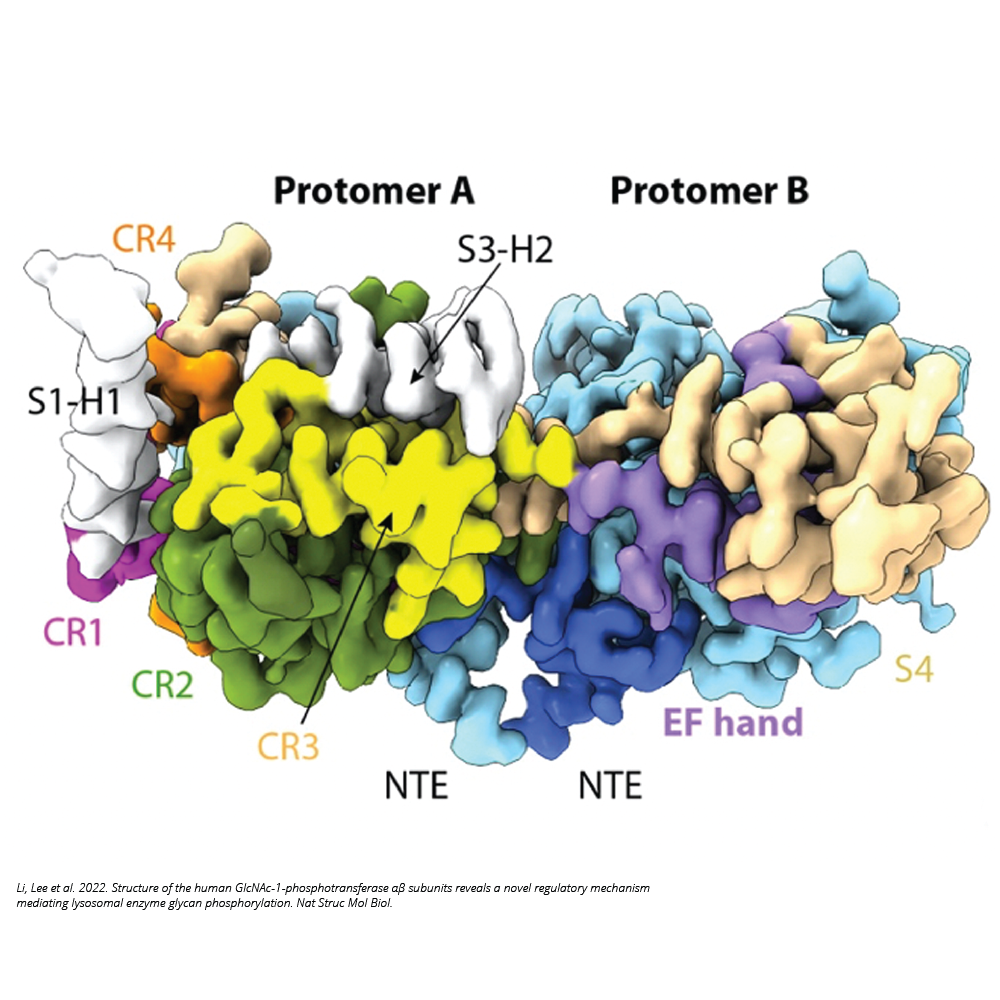
Glycans on proteins can be further modified for new functions. For example, vertebrates have evolved the mannose 6-phosphate (M6P) recognition system to deliver lysosomal hydrolases to lysosomes. In this system, a phosphotransferase (PTase, N-acetylglucosamine (GlcNAc)-1-phosphotransferase) adds GlcNAc-phosphate to mannose residues in the N-glycan of the lysosomal hydrolases. We have determined the structure of the human PTase and discovered a hockey stick-like motif that controls activation of the enzyme (Li et al. 2022, Nat Struct Mol Biol). More recently, we found that a highly truncated human PTase is hyperactive because the truncation has removed the hockey stick, making the enzyme in a constitutively active state (Li et al. 2024, J Biol Chem). Interestingly, despite the truncated PTase resembles the simpler PTase homolog in fruit fly, the fly enzyme does not act on lysosomal hydrolases (Li et al. 2024, J Biol Chem).
Concomitant to their glycosylation, nascent membrane proteins are inserted into the lipid bilayer with the help of membrane chaperones such as the ER membrane complex whose structure was determined by us recently (Bai et al. 2020, Nature; Bai et al. 2023, Curr Opin Struct Biol). Then, the glycosylated and folded membrane proteins are trafficked from ER through Golgi to reach cell surface membrane. This transport process is facilitated by several lipid flippases. We have determined the structure and substrate binding and flipping mechanisms for three yeast lipid flippases: Drs2-Cdc50 (Bai et al. 2019, Nat Commun), Neo1 (Bai et al. 2021, Nat Commun), and Dnf1/Dnf2-Cdc50 (Bai et al. 2020, eLife). This work has improved our understanding of molecular mechanisms of flipping lipids from outer leaflet to the inner leaflet of the membrane bilayers (Duan et al. 2024, J Biol Chem). To further understand how the Drs2-Cdc50 flippase is activated by Arl1-Gea2 complex, we have recently revealed that Gea2 assembles a stable dimer, and unexpectedly, Arl1 binds to the DCB rather than the SEC7 domain (Duan et al. 2024, Nat Commun).
Related Publications
Bai L, Wang T, Zhao G, Kovach A, Li H. 2018. The atomic structure of a eukaryotic oligosaccharyltransferase complex. Nature 555(7696):328–333.
Bai L, Kovach A, You Q, Hsu C, Zhao G, Li H. 2019. Autoinhibition and activation mechanisms of the eurkaryotic lipid flippase Drs2p-Cdc50p. Nat Commun 10(1):4142.
Bai L, Kovach A, You Q, Kenny A, Li H. 2019. Structure of the eukaryotic protein O-mannosyltransferase Pmt1-Pmt2 complex. Nat Struct Mol Biol 26(8):704–711.
Bai L, You Q, Feng X, Kovach A, Li H. 2020. Structure of the ER membrane complex, a transmembrane-domain insertase. Nature 584:475–478.
Bai L, Kovach A, You Q, Hsu C, Zhao G, Li H. 2019. Autoinhibition and activation mechanisms of the eurkaryotic lipid flippase Drs2p-Cdc50p. Nat Commun 10(1):4142.
Bai L, You Q, Jain BK, Duan HD, Kovach A, Graham TR, Li H. 2020. Transport mechanism of P4 ATPase phosphatidylcholine flippases. eLife 9:e62163.
Bai L, Jain BK, You Q, Duan HD, Takar M, Graham TR, Li H. 2021. Structural basis of the P4B ATPase lipid flippase activity. Nat Commun 12:5963.
Bai L, Li H. 2023. Structural insights into the membrane chaperones for multi-pass membrane protein biogenesis. Curr Opin Struct Biol 79:102563.
Duan HD, Li H. 2024. Consensus, controversies, and conundrums of P4-ATPases: the emerging face of eukaryotic lipid flippases. J Biol Chem 300(6):107387.
Duan HD, Jain BK, Li H, Graham TR*, Li H*. 2024. Structural insight into an Arl1-ArfGEF complex involved in Golgi recruitment of a GRIP-domain golgin. Nat Commun 15:1942.
*Co-corresponding authors
Harada Y, Li H, Li H, Lennarz WJ. 2009. Oligosaccharyltransferase directly binds to ribosome at a location near the translocon-binding site. Proc Natl Acad Sci U S A 106(17):6945–6949.
Li H, Chapla D, Amos RA, Ramiah A, Moremen KW, Li H. 2023. Structural basis for heparan sulfate co-polymerase action by the EXT1–2 complex. Nat Chem Biol.
Li H, Chavan M, Schindelin H, Lennarz WJ, Li H. 2008. Structure of the oligosaccharyl transferase complex at 12 Å resolution. Structure 16(3):432–440.
Li H, Doray B, Jennings BC, Lee W-S, Liu L, Kornfeld S, Li H. 2024. Structure of a truncated human GlcNAc-1-phosphotransferase variant reveals the basis for its hyperactivity. J Biol Chem 300(9):107706.
Li H, Lee WS, Feng X, Bai L, Jennings BC, Liu L, Doray B, Canfield WM, Kornfeld S, Li H. 2022. Structure of the human GlcNAc-1-phosphotransferase αβ subunits reveals regulatory mechanism for lysosomal enzyme glycan phosphorylation. Nat Struct Molec Biol 29(4):348–356.
Takeuchi H, Yu H, Hao H, Takeuchi M, Ito A, Li H, Haltiwanger RS. 2017. O-Glycosylation modulates the stability of epidermal growth factor-like repeats and thereby regulates Notch trafficking. J Biol Chem 292(38):15964–15973.
Yu H, Takeuchi M, LeBarron J, Kantharia J, London E, Bakker H, Haltiwanger RS, Li H*, Takeuchi H*. 2015. Notch-modifying xylosyltransferase structures support an SNi-like retaining mechanism. Nat Chem Biol 11(11):847–854.
Yu H, Takeuchi H, Takeuchi M, Liu Q, Kantharia J, Haltiwanger RS, Li H. 2016. Structural analysis of Notch-regulating Rumi reveals basis for pathogenic mutations. Nat Chem Biol 12(9):735–740.
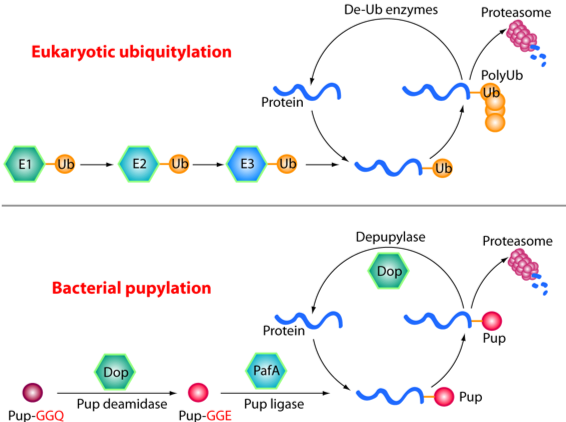
The Pup-proteasome System in Mycobacterium Tuberculosis
Tuberculosis (TB) kills 1.5 million to 2 million people globally every year. In the U.S. alone, 13 million people have latent TB infection; and about 10% of them will develop active TB disease during their lifetime. An effective vaccine or chemotherapy has yet to be developed. Mycobacterium tuberculosis (Mtb) — the bacterium that causes TB — has evolved a Pup-proteasome system that is functionally analogous to but chemically distinct from the eukaryotic Ubiquitin-proteasome system. The Pup-proteasome is required for Mtb resistance to killing by a source of nitric oxide (NO). NO is required by the host immune system to control Mtb infections. Thus, the Pup-proteasome system is a promising target for the development of anti-TB chemotherapeutics.
In collaboration with the labs of Dr. Carl Nathan, Dr. Gang Lin and Dr. Heran Darwin, we have combined cryo-EM, X-ray crystallography and protein biochemistry to elucidate the structure and function of the bacterial system. We found that the Mtb proteasome is structurally similar to the eukaryotic proteasome yet possesses unique assembly and gating mechanisms (Hu et al. 2006, Mol Microbiol; Li et al. 2010, EMBO J). Like its eukaryotic counterpart, the bacterial proteasome can be activated either by an ATP-dependent activator (Mpa) (Wu et al. 2017, Mol Microbiol; Yin et al. 2021, J Biol Chem), or by an ATP-independent activator (PafE) (Jastrab et al. 2015, Proc Natl Acad Sci U S A; Bai et al. 2016, Proc Natl Acad Sci U S A; Hu et al. 2018, J Biol Chem). Interestingly, we found that the protein degradation tag Pup, a prokaryotic Ubiquitin-like protein, is intrinsically disordered, but folds into an α-helix upon binding to and recognized by the proteasomal ATPase Mpa (Wang et al. 2010, Nat Struct Mol Biol). To facilitate the development of anti-TB chemotherapeutics, we have elucidated the structural basis for species-specific inhibition of the Mtb proteasome by oxathiazol-2-ones (Lin et al. 2009, Nature), by several N,C-capped dipeptides (Hsu et al. 2017, Biochemistry), and by the phenylimidazo-based and macrocyclic peptide-based inhibitors (Zhan et al. 2019, J Med Chem; Zhang et al. 2021, J Med Chem). Remarkably, some of the dipeptide variants exhibited potent and selective inhibition to the human immunoproteasome (Santos et al. 2017, Nat Commun).
In addition to removing the damaged proteins by the Pup-proteasome system, Mtb also uses the DnaK-ClpB bi-chaperone system to rescue and refold damaged proteins. We have studied the protein disaggregation mechanism by the Mtb ClpB hexamer (Yu et al. 2018, Proc Natl Acad Sci U S A) and revealed how ClpB and DnaK work together to synergistically disaggregate and refold protein aggregates (Yin et al. 2021, Cell Rep), and discovered that the GrpE dimer facilitates not only the nucleotide exchange but also the protein substrate release from the DnaK (Xiao et al. 2024, Nat Commun).
Related Publications
Bai L, Hu K, Wang T, Jastrab JB, Darwin KH, Li H. 2016. Structural analysis of the dodecameric proteasome activator PafE in Mycobacterium tuberculosis. Proc Natl Acad Sci U S A 113(14):E1983–1992.
Hu G, Lin G, Wang M, Dick L, Xu RM, Nathan C, Li H. 2006. Structure of the Mycobacterium tuberculosis proteasome and mechanism of inhibition by a peptidyl boronate. Mol Microbiol 59(5):1417–1428.
Hsu HC, Singh PK, Fan H, Wang R, Sukenick G, Nathan C, Lin G, Li H. 2017. Structural basis for the species-selective binding of N,C-capped dipeptides to the Mycobacterium tuberculosis proteasome. Biochemistry 56(1):324–333.
Jastrab JB, Wang T, Murphy JP, Bai L, Hu K, Merkx R, Huang J, Chatterjee C, Ovaa H, Gygi SP, Li H, Darwin KH. 2015. An adenosine triphosphate-independent proteasome activator contributes to the virulence of Mycobacterium tuberculosis. Proc Natl Acad Sci U S A. 112(14):E1763–1772.
Li D, Li H, Wang T, Pan H, Lin G, Li H. 2010. Structural basis for the assembly and gate closure mechanisms of the Mycobacterium tuberculosis 20S proteasome. EMBO J. 29(12):2037–2047.
Lin G*, Li D, de Carvalho L, Deng H, Tao H, Vogt G, Wu K, Schneider J, Chidawanyika T, Warren JD, Li H*, Nathan C*. 2009. Inhibitors selective for mycobacterial versus human proteasomes. Nature 461(7264):621–626.
Santos RLA, Bai L, Singh PK, Murakami N, Fan H, Zhan W, Zhu Y, Jiang X, Zhang K, Assker JP, Nathan CF, Li H, Azzi J, Lin G. 2017. Structure of human immunoproteasome with a reversible and noncompetitive inhibitor that selectively inhibits activated lymphocytes. Nat Commun 8(1):1692.
Hu K, Jastrab JB, Zhang S, Kovach A, Zhao G, Darwin KH, Li H. 2018. Proteasome substrate capture and gate opening by the accessory factor PafE from Mycobacterium tuberculosis. J Biol Chem 293(13):4713–4723.
Samanovic MI, Hsu HC, Jones MB, Jones V, McNeil MR, Becker SH, Jordan AT, Strnad M, Xu C, Jackson M, Li H, Darwin KH. 2018. Cytokinin signaling in Mycobacterium tuberculosis. MBio 19(3).
Wang T, Darwin KH, Li H. 2010. Binding-induced folding of prokaryotic ubiquitin-like protein (Pup) targets substrates for proteasomal degradation. Nat Struct Mol Biol 17(11):1352–1357.
Wu Y, Hu K, Li D, Bai L, Yang S, Jastrab JB, Xiao S, Hu Y, Zhang S, Darwin KH, Wang T, Li H. 2017. Mycobacterium tuberculosis proteasomal ATPase Mpa has a β-grasp domain that hinders docking with the proteasome core protease. Mol Microbiol 105(2):227–241.
Xiao X, Fay A, Santos Molina P, Kovach A, Glickman MS, Li H. 2024. Structure of the M. tuberculosis DnaK−GrpE complex reveals how key DnaK roles are controlled. Nat Comm 15:660.
Yin Y, Kovach A, Hsu HC, Darwin KH, Li H. 2021. The mycobacterial proteasomal ATPase Mpa forms a gapped ring to engage the 20S proteasome. J Biol Chem 100713.
Yin Y, Feng X, Yu H, Fay A, Kovach A, Glickman MS, Li H. 2021. Structural basis for aggregate dissolution and refolding by the Mycobacterium tuberculosis ClpB-DnaK bi-chaperone system. Cell Rep 35(8):109166.
Yu H, Lupoli TJ, Kovach A, Meng X, Zhao G, Nathan CF, Li H. 2018. ATP hydrolysis-coupled peptide translocation mechanism of Mycobacterium tuberculosis ClpB. Proc Natl Acad Sci U S A 115(41)::E9560–E9569.
Zhan W, Hsu HC, Morgan T, Oullette T, Burns-Huang K, Hara R, Wright AG, Imaeda T, Okamoto R, Sato K, Michino M, Ramjee M, Aso K, Meinke P, Foley M, Nathan CR, Li H, Lin G. 2019. Selective phenylimidazole-based inhibitors of the Mycobacterium tuberculosis proteasome. J Med Chem 62(20):9246–9253.
Zhang H, Hsu HC, Kahne SC, Hara R, Zhan W, Jiang X, Burns-Huang K, Ouellette T, Imaeda T, Okamoto R, Kawasaki M, Michino M, Wong TT, Toita A, Yukawa T, Moraca F, Vendome J, Saha P, Sato K, Aso K, Ginn J, Meinke PT, Foley M, Nathan CF, Darwin KH, Li H, Lin G. 2021. Macrocyclic peptides that selectively inhibit the Mycobacterium tuberculosis proteasome. J Med Chem 64(9):6262–6272.
Molecular Mechanisms of Energy Conservation
Energy conservation in modern-day mitochondria relies on oxygen as the ultimate electron acceptor. However, life evolved 4 billion years ago, and atmospheric oxygen was available only 2.5 billion years ago. In the absence of oxygen, ancestral life has used proton or elemental sulfur as electron acceptors, mediated by the ancestral respiratory complexes such as the membrane-bound hydrogenase (MBH) and membrane-bound sulfane sulfur reductase (MBS), which now exist in hyperthermophile Pyrococcus furiosus. We have recently solved the structures of MBH (Yu et al. 2018, Cell) and MBS (Yu et al. 2021, Nat Commun), shedding light on their redox-coupled ion pumping mechanisms and their evolutionary relationships with the modern-day respiratory Complex I (Yu et al. 2021, JBC review).
Electron bifurcation was first discovered by Peter Mitchell in respiratory Complex III and is quinone-based. Recently, flavin-based electron bifurcation (FBEB) is found to be a widespread energy conservation mechanism in microbial metabolism. These oxidoreductase-type enzymes catalyze endergonic reduction of low-potential, high-energy substrates by simultaneously coupling them to the reduction of high-potential, low-energy substrates in an exergonic reaction. Electron bifurcation is therefore used by microorganisms as a general mechanism to produce low-potential, high-energy electrons without the expenditure of ATP. In collaboration with the Michael Adams lab, we have recently solved the structures of the membrane-associated Fix/EtfABCX (Feng et al. 2021, Proc Natl Acad Sci U S A) and the NiFe-HydABCSL (Feng et al. 2022, Sci Adv, in print), revealing their electron transport pathways and potential bifurcation mechanisms.
Related Publications
Feng X, Schut G, Lipscomb G, Li H, Adams M. 2021. Cryoelectron microscopy structure and mechanism of the membrane-associated electron-bifurcating flavoprotein Fix/EtfABCX. Proc Natl Acad Sci U S A 118(2).
Feng X, Schut G, Haja D, Adama M, Li H. 2022. Structure and electron transfer pathways of an electron bifurcating NiFe-hydrogenase. Sci Adv.
Yu H, Wu CH, Schut G, Haja D, Zhao G, Peters J, Adams M, Li H. 2018. Structure of an ancient respiratory system. Cell 173(7):1636-1649.
Yu H, Haja DK, Schut GJ, Wu CH, Meng X, Zhao G, Li H, Adams MWW. 2020. Structure of the respiratory MBS complex reveals iron-sulfur cluster catalyzed sulfane sulfur reduction in ancient life. Nat Commun 11(1):5953.
Yu H, Schut G, Haja D, Adams M, Li H. 2021. Evolution of complex I-like respiratory complexes. J Biol Chem 296: 100740. (Invited review)
SELECTED PUBLICATIONS
For a full list of Dr. Li’s publications, please visit his NCBI bibliography.
Feng X, Spiering MM, Ruda de Luna Santos A, Benkovic SJ, Li H. 2025. Structural insights into the exchange mechanism of a replicative DNA polymerase. Nucleic Acids Res 53(22).
Du M, Yuan Z, Kovach A, Lyu M, Li H. 2025. Pmt4 recognizes two separate acceptor sites to O-mannosylate in the S/T rich regions of substrate proteins. Nat Commun 16(1).
*Selected as a featured paper
Jain BK, Duan HD, Valentine C, Samiha A, Li H, Graham TR. 2025. P4-ATPases control phosphoinositide membrane asymmetry and neomycin resistance. Nat Cell Biol 27(7):114-1124.
Xiao X, Schut G, Feng X, McTernan PM, Haja DK, Lanzilotta WN, Adams MWW, Li H. 2025. Structural insights into the biotechnologically relevant reversible NADPH-oxidizing NiFe-hydrogenase from P. furiosus. Structure.
Wang F, He Q, O’Donnell ME, Li H. 2025. The proofreading mechanism of the human leading-strand DNA polymerase holoenzyme. Proc Natl Acad Sci U S A 122(22):e2507232122.
Li H, Doray B, Jennings BC, Lee W-S, Liu L, Kornfeld S, Li H. 2024. Structure of a truncated human GlcNAc-1-phosphotransferase variant reveals the basis for its hyperactivity. J Biol Chem 300(9):107706.
He Q, Wang F, Yao NY, O’Donnell ME*, Li H*. 2024. Structures of the human leading strand Polε–PCNA holoenzyme. Nat Commun 15(7847).
*Co-corresponding authors
Yuan Z, Georgescu R, Yao NY, Yurieva O, O’Donnell ME, Li H. 2024. Mechanism of PCNA loading by Ctf18-RFC for leading-strand DNA synthesis. Science 385(6708).
Feng X, Schut GJ, Adams MWW, Li H. Structures and electron transport paths in the four families of flavin-based electron bifurcation enzymes. In: Harris JR, Marles-Wright J (eds). Macromolecular Protein Complexes V: Structure and Function, Springer-Nature: 383-408.
Duan HD, Li H. 2024. Consensus, controversies, and conundrums of P4-ATPases: the emerging face of eukaryotic lipid flippases. J Biol Chem 300(6):107387.
Wang F, He Q, Yao NY, O’Donnell ME, Li H. 2024. The human ATAD5 has evolved unique structural elements to function exclusively as a PCNA unloader. Nat Struct Mol Biol.
Yuan Z, Li H. 2024. Primase and polymerase α tango to make an RNA-DNA hybrid primer. FEBS J 291(9):1889–1891.
He Q, Wang F, O’Donnell ME, Li H. 2024. Cryo-EM reveals a nearly complete PCNA loading process and unique features of the human alternative clamp loader CTF18-RFC. Proc Nat Acad Sci U S A 121(18):e2319727121.
Duan HD, Jain BK, Li H, Graham TR*, Li H*. 2024. Structural insight into an Arl1-ArfGEF complex involved in Golgi recruitment of a GRIP-domain golgin. Nat Commun 15:1942.
*Co-corresponding authors
Zheng F, Yao NY, Georgescu RE, Li H*, O’Donnell ME*. 2024. Structure of the PCNA unloader Elg1-RFC. Sci Adv 10(9).
*Co-corresponding authors
Xiao X, Fay A, Santos Molina P, Kovach A, Glickman MS, Li H. 2024. Structure of the M. tuberculosis DnaK−GrpE complex reveals how key DnaK roles are controlled. Nat Comm 15:660.
Hsu HC, Li D, Zhan W, Ye J, Liu YJ, Leung A, Qin J, Crespo B, Gamo FJ, Zhang H, Cui L, Roth A, Kirkman LA, Li H, Lin G. 2023. Structures revealing mechanisms of resistance and collateral sensitivity of Plasmodium falciparum to proteasome inhibitors. Nat Commun 14(1):8302.
Zheng F, Georgescu RE, Yao NY, O’Donnell ME, Li H. 2023. Structures of 9-1-1 DNA checkpoint clamp loading at gaps from start to finish and ramification on biology. Cell Rep 42(7):112694.
Feng X, Spiering MM, de Luna Almeida Santos R, Benkovic SJ, Li H. 2023. Structural basis of the T4 bacteriophage primosome assembly and primer synthesis. Nat Commun 14(1):4396.
Burgie ES, Li H, Gannam ZTK, McLoughlin KE, Vierstra RD*, Li H*. 2023. The structure of Arabidopsis phytochrome A reveals topological and functional diversification among the plant photoreceptor isoforms. Nat Plants 9(7):1116–1129.
*Co-corresponding authors
Hsu HC, Wang J, Kjellgren A, Li H, DeMartino GN. 2023. Ηigh-resolution structure of mammalian PI31-20S proteasome complex reveals mechanism of proteasome inhibition. J Biol Chem 299(7):104862.
Bai L, Li H. 2023. Structural insights into the membrane chaperones for multi-pass membrane protein biogenesis. Curr Opin Struct Biol 79:102563.
Yuan Z, Georgescu R, Li H*, O’Donnell ME*. 2023. Molecular choreography of primer synthesis by the eukaryotic Pol α-primase. Nat Commun 14(1):3697.
*Co-corresponding authors
Wang F, He Q, Zhan W, Yu Z, Finkin-Groner E, Ma X, Lin G, Li H. 2023. Structure of the human UBR5 E3 ubiquitin ligase. Structure.
Li H, Chapla D, Amos RA, Ramiah A, Moremen KW, Li H. 2023. Structural basis for heparan sulfate co-polymerase action by the EXT1–2 complex. Nat Chem Biol.
Bai L, Li H. 2022. Cryo-EM structures of the endoplasmic reticulum membrane complex. FEBS J 289(1):102–112.
Langston LD, Yuan Z, Georgescu R, Li H, O’Donnell ME. 2022. SV40 T-antigen uses a DNA shearing mechanism to initiate origin unwinding. Proc Natl Acad Sci U S A 119(49):e2216240119.
Li H, Lee WS, Feng X, Bai L, Jennings BC, Liu L, Doray B, Canfield WM, Kornfeld S, Li H. 2022. Structure of the human GlcNAc-1-phosphotransferase αβ subunits reveals regulatory mechanism for lysosomal enzyme glycan phosphorylation. Nature Struct Molec Biol 29(4):348–356.
Li H, O’Donnell M, Kelch B. 2022. Unexpected new insights into DNA clamp loaders. Bioessays 44(11):e2200154.
Schut GJ, Haja DK, Feng X, Poole FL, II, Li H, Adams MWW. 2022. An abundant and diverse new family of electron bifurcating enzymes with a non-canonical catalytic mechanism. Front Microbiol 13:946711.
Wang F, Feng X, He Q, Li H, Li H. 2022. The S. Cerevisiae Yta7 ATPase hexamer contains a unique bromodomain tier that functions in nucleosome disassembly. J Biol Chem 102852.
Xiao X, Feng X, Yoo JH, Kovach A, Darwin KH, Li H. 2022. The β-grasp domain of proteasomal ATPase Mpa makes critical contacts with the Mycobacterium tuberculosis 20S core particle to facilitate degradation. mSphere 7(5):e00227422.
Zheng F, Georgescu R, Yao NY, Li H, O’Donnell ME. 2022. Cryo-EM structures reveal that RFC recognizes both the 3′- and 5′-DNA ends to load PCNA onto gaps for DNA repair. eLife 11:e77469.
Li H*, Burgie S*, Gannam ZTK, Li H#, Vierstra RD#. 2022. Plant phytochromes B is an asymmetric dimers with unique signaling potential. Nature 604(7904): 127-133.
* Co-first authors
# Co-corresponding authors
Li H#, Lee WS#, Feng X, Bai L, Jennings JC, Liu L, Doray B, Canfield WM, Kornfeld S*, Li H*. 2022. Structure of the human GlcNAc-1-phosphotransferase αβ subunits reveals regulatory mechanism for lysosomal enzyme glycan phosphorylation. Nat Struct Mol Biol 29(4): 348-356.
# Co-first authors
* Co-corresponding authors
Zheng F, Georgescu RE, Yao NY, O’Donnell ME*, Li H*. 2022. DNA is loaded through the 911 DNA checkpoint clamp in the opposite direction of the PCNA clamp. Nat Struc Mol Biol 29(4): 378-385.
* Co-corresponding authors
Feng X, Schut G, Haja D, Adama M, Li H. 2022. Structure and electron transfer pathways of an electron bifurcating NiFe-hydrogenase. Sci Adv 8(8): eabm7546.
Hsu HC, Wang M, Kovach A, Darwin AJ, Li H. 2022. Pseudomonas aeruginosa C-terminal processing protease CtpA assembles into a hexameric structure that requires activation by a spiral-shaped lipoprotein-binding partner. mBio 13(1): e0368021.
Zhao P, Zhao C, Chen D, Yun C, Li H*, Bai L*. 2021. Structure and activation mechanism of the hexameric plasma membrane H+-ATPase. Nat Commun 12:6439.
*Co-corresponding authors
Included as an Editor’s Highlight
Bai L, Jain BK, You Q, Duan HD, Takar M, Graham TR, Li H. 2021. Structural basis of the P4B ATPase lipid flippase activity. Nat Commun 12:5963.
Du M, Yuan Z, Werneburg GT, Henderson NS, Chauhan H, Kovach A, Zhao G, Johl J, Li H, Thanassi DG. 2021. Processive dynamics of the usher assembly platform during uropathogenic Escherichia coli P pilus biogenesis. Nat Commun 12(1):5207.
Yu H, Schut GJ, Haja DK, Adams MWW, Li H. 2021. Evolution of complex I-like respiratory complexes. JBC Rev 296:100740.
Yin Y, Feng X, Yu H, Fay A, Kovach A, Glickman MS, Li H. 2021. Structural basis for aggregate dissolution and refolding by the Mycobacterium tuberculosis ClpB-DnaK bi-chaperone system. Cell Rep 35(8):109166.
Zhang H, Hsu HC, Kahne SC, Hara R, Zhan W, Jiang X, Burns-Huang K, Ouellette T, Imaeda T, Okamoto R, Kawasaki M, Michino M, Wong TT, Toita A, Yukawa T, Moraca F, Vendome J, Saha P, Sato K, Aso K, Ginn J, Meinke PT, Foley M, Nathan CF, Darwin KH, Li H, Lin G. 2021. Macrocyclic peptides that selectively inhibit the Mycobacterium tuberculosis proteasome. J Med Chem 64(9):6262–6272.
Yin Y, Kovach A, Hsu HC, Darwin KH, Li H. 2021. The mycobacterial proteasomal ATPase Mpa forms a gapped ring to engage the 20S proteasome. J Biol Chem 100713.
Bai L, Li H. 2021. Cryo-EM structures of the endoplasmic reticulum membrane complex. FEBS J.
Li H, Zheng F, O’Donnell M. 2021. Water skating: How polymerase sliding clamps move on DNA. FEBS J.
Feng X, Schut GJ, Lipscomb GL, Li H, Adams MWW. 2021. Cryoelectron microscopy structure and mechanism of the membrane-associated electron-bifurcating flavoprotein Fix/EtfABCX. Proc Natl Acad Sci U S A 118(2).
Bai L, Li H. 2021. Protein N-glycosylation and O-mannosylation are catalyzed by two evolutionarily related GT-C glycosyltransferases. Curr Opin Struct Biol 68:66–73.
Bohl T, Bai L, Li H. 2021. Recent progress in structural studies on the GT-C superfamily of protein glycosyltransferases. Subcell Biochem 96:259–271.
Li H, Yao NY, O’Donnell ME. 2020. Anatomy of a twin DNA replication factory. Biochem Soc Trans 48(6):2769–2778.
Zheng F, Georgescu RE, Li H, O’Donnell ME. 2020. Structure of eukaryotic DNA polymerase δ bound to the PCNA clamp while encircling DNA. Proc Natl Acad Sci U S A 117(48):30344–30353.
Yu H, Haja DK, Schut GJ, Wu CH, Meng X, Zhao G, Li H, Adams MWW. 2020. Structure of the respiratory MBS complex reveals iron-sulfur cluster catalyzed sulfane sulfur reduction in ancient life. Nat Commun 11(1):5953.
Yuan Z, Li H. 2020. Molecular mechanisms of eukaryotic origin initiation, replication fork progression, and chromatin maintenance. Biochem J 477(18):3499–3525.
Yuan Z, Schneider S, Dodd T, Riera A, Bai L, Yan C, Magdalou I, Ivanov I, Stillman B, Li H, Speck C. 2020. Structural mechanism of helicase loading onto replication origin DNA by ORC-Cdc6. Proc Natl Acad Sci U S A 117(30):17747–17756.
Yuan Z, Georgescu R, Schauer GD, O’Donnell ME, Li H. 2020. Structure of the polymerase ε holoenzyme and atomic model of the leading strand replisome. Nat Commun 11(1):3156.
Yuan Z, Georgescu R, Bai L, Zhang D, Li H, O’Donnell ME. 2020. DNA unwinding mechanism of a eukaryotic replicative CMG helicase. Nat Commun 11(1):688.
Bai L, You Q, Jain BK, Duan HD, Kovach A, Graham TR, Li H. 2020. Transport mechanism of P4 ATPase phosphatidylcholine flippases. eLife.
Bai L, You Q, Feng X, Kovach A, Li H. 2020. Structure of the ER membrane complex, a transmembrane-domain insertase. Nature.
Yuan Z, Georgescu R, Santos RLA, Zhang D, Bai L, Yao NY, Zhao G, O’Donnell ME, Li H. 2019. Ctf4 organizes sister replisomes and Pol α into a replication factory. eLife e47405.
Zhan W, Hsu HC, Morgan T, Oullette T, Burns-Huang K, Hara R, Wright AG, Imaeda T, Okamoto R, Sato K, Michino M, Ramjee M, Aso K, Meinke P, Foley M, Nathan CR, Li H, Lin G. 2019. Selective phenylimidazole-based inhibitors of the Mycobacterium tuberculosis proteasome. J Med Chem.
Bai L, Kovach A, You Q, Hsu C, Zhao G, Li H. 2019. Autoinhibition and activation mechanisms of the eurkaryotic lipid flippase Drs2p-Cdc50p. Nat Commun 10(1):4142.
Li H, O’Donnell ME. 2019. DNA replication from two different worlds. Science 363(6429):814–815.
Bai L, Kovach A, You Q, Kenny A, Li H. 2019. Structure of the eukaryotic protein O-mannosyltransferase Pmt1−Pmt2 complex. Nat Struct Mol Biol.
Bai L, Li H. 2019. Cryo-EM is uncovering the mechanism of eukaryotic protein N-glycosylation. FEBS 286(9):1638–1644.
Hu K, Jordan AT, Zhang S, Dhabaria A, Kovach A, Rangel M, Ueberheide B, Li H, Darwin KH. 2019. Characterization of guided entry of tail-anchored proteins 3 homologues in Mycobacterium tuberculosis. J Bacteriol.
Samanovic MI, Hsu HC, Jones MB, Jones V, McNeil MR, Becker SH, Jordan AT, Strnad M, Xu C, Jackson M, Li H, Darwin KH. 2018. Cytokinin signaling in Mycobacterium tuberculosis. MBio 19(3).
O’Donnell ME, Li H. 2018. The ring-shaped hexameric helicases that function at DNA replication forks. Nat Struct Mol Biol 25(2):122–130.
Du M, Yaun Z, Yu H, Henderson N, Sarowar S, Zhao G, Werneburg GT, Thanassi DG, Li H. 2018. Handover mechanism of the growing pilus by the bacterial outer-membrane usher FimD. Nature.
Yu H, Wu CH, Schut GJ, Haja DK, Zhao G, Peters JW, Adams MWW, Li H. 2018. Structure of an ancient respiratory system. Cell.
Li H, O’Donnell ME. 2018. The eukaryotic CMG helicase at the replication fork: Emerging architecture reveals an unexpected mechanism. BioEssays40(3):1700208.
*Highlighted in Advanced Science News.
Bai L, Wang T, Zhao G, Kovach A, Li H. 2018. The atomic structure of a eukaryotic oligosaccharyltransferase complex. Nature.
Bai L, Yuan Z, Sun J, Georgescu R, O’Donnell ME, Li H. 2017. Architecture of the Saccharomyces cerevisiae replisome. Adv Exp Med Biol 1042:207–228.
Noguchi Y, Yuan Z, Bai L, Schneider S, Zhao G, Stillman B, Speck C, Li H. 2017. Cryo-EM structure of Mcm2-7 double hexamer on DNA suggests a lagging-strand DNA extrusion model. Proc Natl Acad Sci U S A.
Wu Y, Hu K, Li D, Bai L, Yang S, Jastrab JB, Xiao S, Hu Y, Zhang S, Darwin KH, Wang T, Li H. 2017. Mycobacterium tuberculosis proteasomal ATPase Mpa has a β-grasp domain that hinders docking with the proteasome core protease. Mol Microbiol 105(2):227–241.
Yuan Z, Riera A, Bai L, Sun J, Nandi S, Spanos C, Chen ZA, Barbon M, Rappsilber J, Stillman B, Speck C, Li H. 2017. Structural basis of MCM2-7 replicative helicase loading by ORC-Cdc6 and Cdt1. Nat Struct Mol Biol 24(3):316–324.
Georgescu R, Yuan Z, Bai L, de Luna Almeida Santos R, Sun J, Zhang D, Yurieva O, Li H, O’Donnell ME. 2017. Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation. Proc Natl Acad Sci U S A 114(5):E697–E706.
Yu H, Takeuchi H, Takeuchi M, Liu Q, Kantharia J, Haltiwanger RS, Li H. 2016. Structural analysis of Notch-regulating Rumi reveals basis for pathogenic mutations. Nat Chem Biol 12:735–740.
Yuan Z, Bai L, Sun J, Georgescu R, O’Donnell ME, Li H. 2016. Structure of the eukaryotic replicative CMG helicase suggests a pumpjack motion for translocation. Nat Struct Mol Biol 23(3):217–224.
O’Donnell M, Li H. 2016. The eukaryotic replisome goes under the microscope. Curr Biol 26(6):R247-56.
Bai L, Hu K, Wang T, Jastrab JB, Darwin KH, Li H. 2016. Structural analysis of the dodecameric proteasome activator PafE in Mycobacterium tuberculosis. Proc Natl Acad Sci U S A 113(14):E1983-92.
Sun J, Shi Y, Georgescu RE, Yuan Z, Chait BT, Li H, O’Donnell ME. 2015. The architecture of a eukaryotic replisome. Nat Struct Mol Biol 22:976–982.
Yu H, Takeuchi M, LeBarron J, Kantharia J, London E, Bakker H, Haltiwanger RS, Li H, Takeuchi H. 2015. Notch-modifying xylosyltransferase structures support an SNi-like retaining mechanism. Nat Chem Biol 11(11):847–854.
Sun J, Fernandez-Cid A, Riera A, Tognetti S, Yuan Z, Stillman B, Speck C, Li H. 2014. Structural and mechanistic insights into licensing of DNA replication. Genes Dev 28:2291–303.
Sun J, Evrin C, Samel S, Fernadez-Cid A, Riera A, Kawakami H, Zech J, Stillman B, Speck C, Li H. 2013. Cryo-EM structure of a helicase loading intermediate containing ORC-Cdc6-Cdt1-MCM2-7 bound to DNA. Nat Struct Mol Biol 20(8):944–951.
Li H, Stillman B. 2012. The origin recognition complex: a biochemical and structural view. Subcell Biochem 62:37–58.
Sun J, Li H. 2010. How to operate a cryo-electron microscope. Methods Enzymol 481:231-49.
Li D, Li Hua, Wang T, Lin G, Li H. 2010. Structural basis for the assembly and gate closure mechanisms of the Mycobacterium tuberculosis 20S proteasome. EMBO J 29:2037–2047.
Wang T, Darwin KH, Li H. 2010. Binding-induced folding of prokaryotic ubiquitin-like protein (Pup) targets substrates for proteasomal degradation. Nat Struct and Mol Biol 17:1352–1357.
Lin G, Li D, de Carvalho L, Deng H, Tao H, Vogt G, Wu K, Schneider J, Chidawanyika T, Warren JD, Li H, and Nathan C. 2009. Inhibitors selective for mycobacterial versus human proteasomes. Nature 461:621–626.


H. Diessel Duan, Ph.D.
Research Scientist, Department of Structural Biology

Nancy Duchaine
Senior Administrative Assistant I, Department of Structural Biology

Qing He, Ph.D.
Postdoctoral Fellow, Li Laboratory
Structural biology of DNA replication and repair

Hao-Chi Hsu, Ph.D.
Research Scientist, Department of Structural Biology

Sofia Ievleva
Ph.D. Student, VAI Graduate School
Thesis project title to be determined

Amanda Kovach, B.S.
Lab Manager, Department of Structural Biology


Meinan Lyu, Ph.D.
Research Scientist, Department of Structural Biology

Chang Sun, Ph.D.
Research Scientist, Department of Structural Biology

Feng Wang, Ph.D.
Research Scientist, Department of Structural Biology

Merissa (Xiansha) Xiao, Ph.D.
Research Scientist, Li Laboratory
Proteasome and protein folding system involved in multi-drug resistance in Mycobacteria

Qinglong You, Ph.D.
Research Scientist, Department of Structural Biology
Epigenetic mechanisms in DNA damage point

Fengwei Zheng, Ph.D.
Research Scientist, Department of Structural Biology
DNA replication and repair


